Biasing the generation of unweighted partonic events at LO
Since v2.5.0, MadGraph5_aMC@NLO can bias the generation of LO unweighted partonic events, using existing or customized biasing weights. This wiki page will cover the details of how to setup and write your own bias function.
The motivation for applying a bias function falls in either of the following two categories:
- a)
It can be employed in order to produce smoother distributions for the tail of a particular observable. I that case the bias function should not impact the physical results obtained, but rather only affect the distribution of the events generated or, in other words, the phase-space sampling. In this case the events will be first generated and unweighted in presence of this biasing weight. MG5aMC will then reweight these unweighted events by their corresponding bias weight, so as to restore the original physical distributions. The resulting events made available to the user will therefore be weighted in this case, and distributed in a manner that enhances the statistics in the desired region of phase-space.
Notice that one can also use the bias function in this way and with a bias weight equal to unity, if something specific must be done with each event, for example on-the-flight plotting.
- b)
It can be used when one wants to modify the integrand so as to really impact the physical results. In this case MG5aMC will generate and unweight events in presence of the bias function, and the resulting unweighted events will be handed over to the user as such. Consequently, the produced events remain unweighted, with a physical distribution impacted by the bias. In this case, the bias function should be understood as a direct modification of the underlying matrix element weight.
This category of bias can be useful for a plethora of applications: ad-hoc unitarisation, accounting for additional merging weights, inclusion of higher order contributions, implementation of non-trivial cuts depending not only on kinematics, etc..
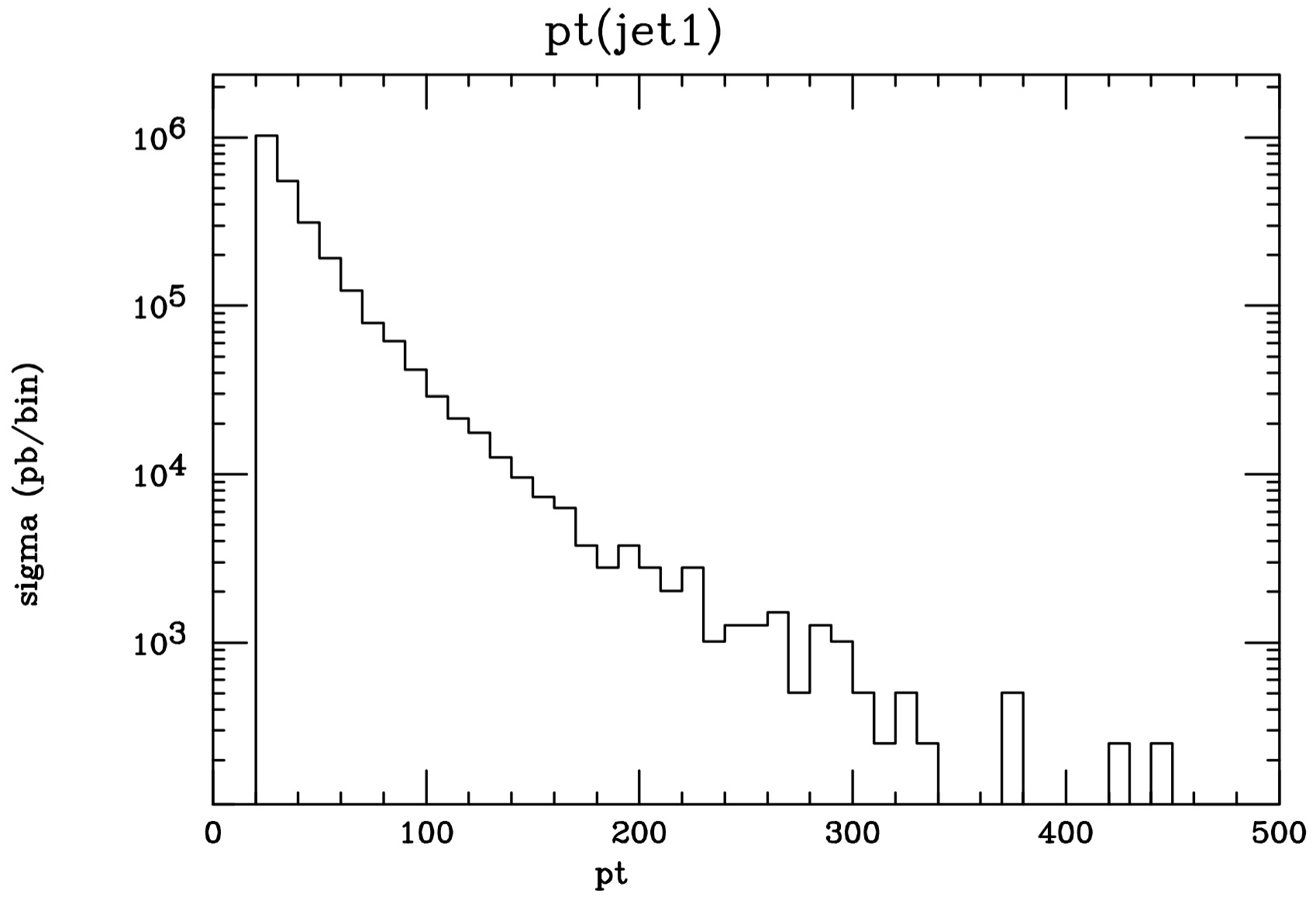
Before going into the detail of the usage of the bias module and the instructions for building your customized bias, we present here an example of its use for the case a) above to smoothen the distribution of the jet transverse momentum for the process 'p p > e+ e- j'.
These two plots were both obtained from 10k events, but for the right-hand side one, the matrix elements have been biased by the quantity $(\;{p_t(j_1)}/{1000.0}\;)4$.
These plots can be straight-forwardly reproduced with (you will need to have install MadAnalysis for the plots to be automatically generated)
./bin/mg5_aMC MG5_aMC> generate p p > e+ e- j MG5_aMC> launch MG5_aMC> set bias_module ptj_bias MG5_aMC> done
Usage of a bias module
In order to activate a given bias module for a MadEvent run, you must specify it in the run card in the entry 'bias_module'.
Alternatively one can use the syntax 'set bias_module <chosen_module>' to write this entry when MG5aMC offers you to edit the cards.
The chosen bias module can either be one of the bias modules natively shipped with MG5aMC ( like 'ptj_bias' ) or an absolute path pointing to a folder that conforms to the format required to implement your own custom bias function (see next section). In any case, in order to turn off any bias, you can set the 'bias_module' run_card entry to 'None'.
Similarly, the bias parameters (all of double precision type) can be specified with a dictionary-like syntax in the run_card entry 'bias_parameters'. For example:
{'ptj_bias_target_ptj': 300.0, 'ptj_bias_enhancement_power': 4.0} = bias_parameters
Alternatively, the commands 'set bias_parameter <param_dictionary>' and 'set bias_parameter <param_name> <value>' can be used when MG5aMC offers you to edit the cards.
Finally, notice that when you choose to use a bias module that does *not* impact the physical results, the unweighted events produced will be re-weighted so as to undo the effect of the bias. As a result the events produced by MadEvent will weighted in this case (obviously unweighting them again would return the original unweighted distribution would would have obtained without the use of a bias in the first place).
Writing your own bias module
In order to develop your custom bias module, you must create a directory that bears the name of the module. For the sake of simplicity, we will illustrate the requirements for a custom bias module with the example of a module that applies on each event a CKKW-L type of merging weight obtained from Pythia8. This bias module module consists in a directory called 'PY8_CKKW' containing the following four files:
- makefile (mandatory)
- PY8_CKKW.f (mandatory)
- bias_dependencies (optional)
- py8_mg5amc_bias_interface.cc (irrelevant for MG5aMC; internal resource of this bias module)
You can also have a look at the structure of the much simpler implementation of the 'ptj_bias' module in '<MG5aMC_distrib_path>/Template/LO/Source/BIAS/ptj_bias'.
The directory 'PY8_CKKW' must then minimally contain the two files 'PY8_CKKW/PY8_CKKW.f' and 'PY8_CKKW/makefile'. The optional file 'PY8_CKKW/bias_dependencies' can also be present.
On top of these files, the user is of course free to include any additional resource that its module will use, but these will not play a role in the interface of the module to MG5_aMC.
Notice however that if you want your module to be working in the cluster mode of MadEvent, you must make sure that your module does not depend on any external file (typically parameter settings that would be read at run-time) since these external files would by default of course not be available on the worker nodes. So you must make sure that everything you will need at run time is already compiled in the library (this is the point of the targets '*_card.h' of the makefile given in example).
Note also that the current bias module only support one multiplicity in the final state so far.
Mandatory file : '<BIAS_MODULE_NAME>/<BIAS_MODULE_NAME>.f'
This fortran file must bear the name of the module with the suffix '.f' and contain the subroutine 'bias_wgt' with the prototype
subroutine bias_wgt(p, original_weight, bias_weight)
include '../../nexternal.inc'
double precision p(0:3,nexternal)
double precision original_weight, bias_weight
Where the array 'p' specifies the kinematics of the events, and the 'original_weight' is also provided, in case your bias function was not to be multiplicative.
The last argument 'bias_weight' is intended as the output.
The parameters of your bias module can be specified by including the following comment in the body of the subroutine:
C parameters = {'OneParam': 0.0,
C 'AnotherParam': 0.0,
C 'YetAnotherOne':2.0}
You must makes sure that your subroutine defines double-precision variables of the same name in its body. The value of these parameters chosen by the user in the run_card can be accessed by including the following include file (after all variable declarations):
include '../bias.inc'
Finally the module must specify the following common block:
double precision stored_bias_weight
data stored_bias_weight/1.0d0/
logical impact_xsec, requires_full_event_info
C Decides whether the bias must be corrected for after unweighting
data impact_xsec/.True./
C Does your bias need the full information about the event
C (color, resonances, helicities, etc..)
data requires_full_event_info/.True./
common/bias/stored_bias_weight,impact_xsec,
& requires_full_event_info
The initialization of the logicals 'impact_xsec' and 'requires_full_event_info' are very important:
- If '
impact_xsec' is set to '.False.', MG5aMC will make sure to reweight the unweighted events produced in presence of the bias so as to undo the physical impact of the bias function (without affecting the resulting event distribution of course). On the contrary, if you want the effect of your bias to persist on the cross-section and differential distribution, you must set this parameter to '.True.'. - If '
requires_full_event_info' is set to '.True.' will make sure to fully process all events sent to your bias module so that color, helicity, resonances information are available. If your module only relies on the kinematics of the event, you should set this parameter to '.False.' so as to speed up the code.
Finally, you should store your desired bias once computed in the subroutine argument 'bias_weight'.
Your module can have access to much more information that just the kinematics specified in the subroutine argument. To access it, simply add the following include file
' include '../../lhe_event_infos.inc'
In particular, one variable defined in this include file is 'character*(maxEventLength) event_record' which stores the full current event record in the LHE format.
There are also other include files that you can add to access more information like the cuts values, the run card parameters, etc...
The illustrative example file 'PY8_CKKW.f' highlights the use of this additional information.
For the sake of completeness, we also show the file 'py8_mg5amc_bias_interface.cc' which implements the C function 'py8_bias_weight' used in 'PY8_CKKW.f'.
Mandatory file : '<BIAS_MODULE_NAME>/makefile'
This makefile will be called by MG5aMC to compile your module into a library when the user choses to use it. It must comply with the following rules:
- a) Include the general makefile options of MG5aMC with:
BIASDIR = ../ SOURCEDIR = ../.. include $(SOURCEDIR)/make_opts
- b) Define the following target '
clean:':
clean: $(RM) *.o $(BIASLIBDIR)$(BIASLIBRARY)
- c) Define the target '
<BIAS_MODULE_NAME>:' that creates the library 'libbias.a' as follows:
<BIAS_MODULE_NAME>: <your list of object.o files> $(call CREATELIB, $(BIASLIBDIR)$(BIASLIBRARY), $^)
- d) Define the target '
requirements:' that prints out 'VALID' if the requirements of your module are met. For example
requirements: ifeq ($(shell $(call CHECK_MG5AMC_VERSION,2.4.2)),True) ifeq ($(PYTHIA8_PATH),NotInstalled) @echo "Error: This module requires Pythia8. Consider installing it with the command 'install pythia8'." else @echo "VALID" endif else @echo "Error:: MG5aMC is not recent enough (found "$(MG5AMC_VERSION)")" @echo "FAIL" endif
which uses the macro 'CHECK_MG5AMC_VERSION' defined in '../make_opts'.
You can then add to this makefile whatever target and dependencies that are necessary for compiling your module.
You can see here the full makefile corresponding to our example module 'PY8_CKKW'.
Optional file : '<BIAS_MODULE_NAME>/bias_dependencies'
Finally your bias module might link against additional libraries that the MadEvent fortran executables should also link to when using the module.
To specify them, you can add the file 'bias_dependencies' to your bias module directory, where you define the variable 'BIASDEPENDENCIES' which will be automatically added to MadEvent makefiles.
Here is what this optional file 'bias_dependencies' looks like in our example module 'PY8_CKKW' (file here):
PY8LIBS = $(shell $(PYTHIA8_PATH)/bin/pythia8-config --libs) PY8INCLUDES = $(shell $(PYTHIA8_PATH)/bin/pythia8-config --includedir) PY8CXXFLAGS = $(shell $(PYTHIA8_PATH)/bin/pythia8-config --cxxflags) BIASDEPENDENCIES = $(PY8LIBS) $(PY8CXXFLAGS)
Note that the environment variable 'PYTHIA8_PATH' will be in this case provided by 'make_opts', also sourced in MadEvent makefiles.
Attachments (7)
- ptj_biased.jpg (114.4 KB ) - added by 10 years ago.
- original_ptj.jpg (111.2 KB ) - added by 10 years ago.
- bias_dependencies (251 bytes ) - added by 10 years ago.
- makefile (2.2 KB ) - added by 10 years ago.
- PY8_CKKW.f (6.0 KB ) - added by 10 years ago.
- PY8_CKKW.2.f (6.0 KB ) - added by 10 years ago.
- py8_mg5amc_bias_interface.cc (4.2 KB ) - added by 10 years ago.
Download all attachments as: .zip